

| Field | Data |
|---|---|
| UniProt Accession: | P11274 |
| Gene Name: | BCR |
| Gene Synonim: | BCR1, D22S11 |
| GAP/GEF: | GAP, GEF |
| Description: | Breakpoint cluster region protein (EC 2.7.11.1) (Renal carcinoma antigen NY-REN-26) |
| Keywords: | 3D-structure;ATP-binding;Acetylation;Alternative splicing;Cell junction;Cell membrane;Chromosomal rearrangement;Complete proteome;GTPase activation;Guanine-nucleotide releasing factor;Kinase;Membrane;Methylation;Nucleotide-binding;Phosphoprotein;Polymorphism;Postsynaptic cell membrane;Proto-oncogene;Reference proteome;Serine/threonine-protein kinase;Synapse;Transferase |
| Entrez ID: | 613 |
| Ensembl Gene ID: | ENSG00000186716 |
| Ensembl Protein ID: | ENSP00000352535 |
| Our Specificity | Rac1 |
| Literature Specificity | RhoA, Rac1 Literature reference |
| Domain Composition: |
 |
| GO ID | Description |
|---|---|
| GO:0005737 | cytoplasm | GO:0043234 | protein complex | GO:0070062 | extracellular exosome | GO:0005829 | cytosol | GO:0016020 | membrane | GO:0045211 | postsynaptic membrane | GO:0014069 | postsynaptic density | GO:0045202 | synapse | GO:0005886 | plasma membrane | GO:0030054 | cell junction |
| GO ID | Description |
|---|---|
| GO:0005089 | Rho guanyl-nucleotide exchange factor activity | GO:0005085 | guanyl-nucleotide exchange factor activity | GO:0016301 | kinase activity | GO:0005096 | GTPase activator activity | GO:0019899 | enzyme binding | GO:0000166 | nucleotide binding | GO:0004674 | protein serine/threonine kinase activity | GO:0016740 | transferase activity | GO:0004713 | protein tyrosine kinase activity | GO:0005515 | protein binding | GO:0005524 | ATP binding |
| GO ID | Description |
|---|---|
| GO:0071222 | cellular response to lipopolysaccharide | GO:0051056 | regulation of small GTPase mediated signal transduction | GO:0060268 | negative regulation of respiratory burst | GO:0016310 | phosphorylation | GO:0060313 | negative regulation of blood vessel remodeling | GO:2000378 | negative regulation of reactive oxygen species metabolic process | GO:0018108 | peptidyl-tyrosine phosphorylation | GO:0048008 | platelet-derived growth factor receptor signaling pathway | GO:0003014 | renal system process | GO:0007165 | signal transduction | GO:0035556 | intracellular signal transduction | GO:0035023 | regulation of Rho protein signal transduction | GO:0032496 | response to lipopolysaccharide | GO:0046777 | protein autophosphorylation | GO:0050766 | positive regulation of phagocytosis | GO:0050728 | negative regulation of inflammatory response | GO:0043547 | positive regulation of GTPase activity | GO:0050885 | neuromuscular process controlling balance | GO:0065002 | intracellular protein transmembrane transport | GO:0030336 | negative regulation of cell migration | GO:0006468 | protein phosphorylation | GO:0030036 | actin cytoskeleton organization | GO:0060216 | definitive hemopoiesis | GO:0048872 | homeostasis of number of cells | GO:0042472 | inner ear morphogenesis | GO:0007420 | brain development | GO:0051726 | regulation of cell cycle | GO:0043314 | negative regulation of neutrophil degranulation | GO:0002692 | negative regulation of cellular extravasation | GO:0043114 | regulation of vascular permeability | GO:0051171 | regulation of nitrogen compound metabolic process |
| Reactome ID | Description |
|---|---|
| R-HSA-5655302 | Signaling by FGFR1 in disease | R-HSA-194840 | Rho GTPase cycle | R-HSA-1839117 | Signaling by cytosolic FGFR1 fusion mutants |
| Kegg ID | Description |
|---|---|
| hsa05200 | Pathways in cancer | hsa05220 | Chronic myeloid leukemia |
| Field | Data |
|---|---|
| Localization in this study: | actin, plasma_membrane |
| Localization in the literature: | plasma membrane Golgi cytosolic |
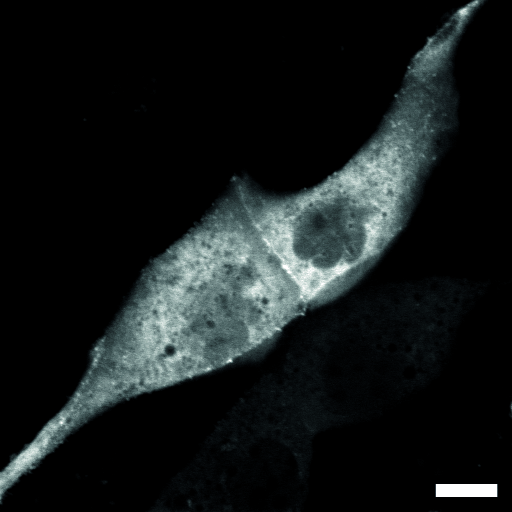
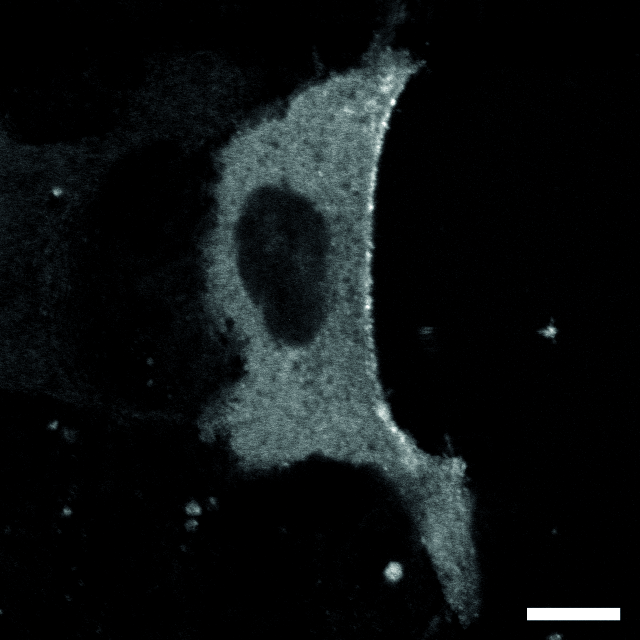
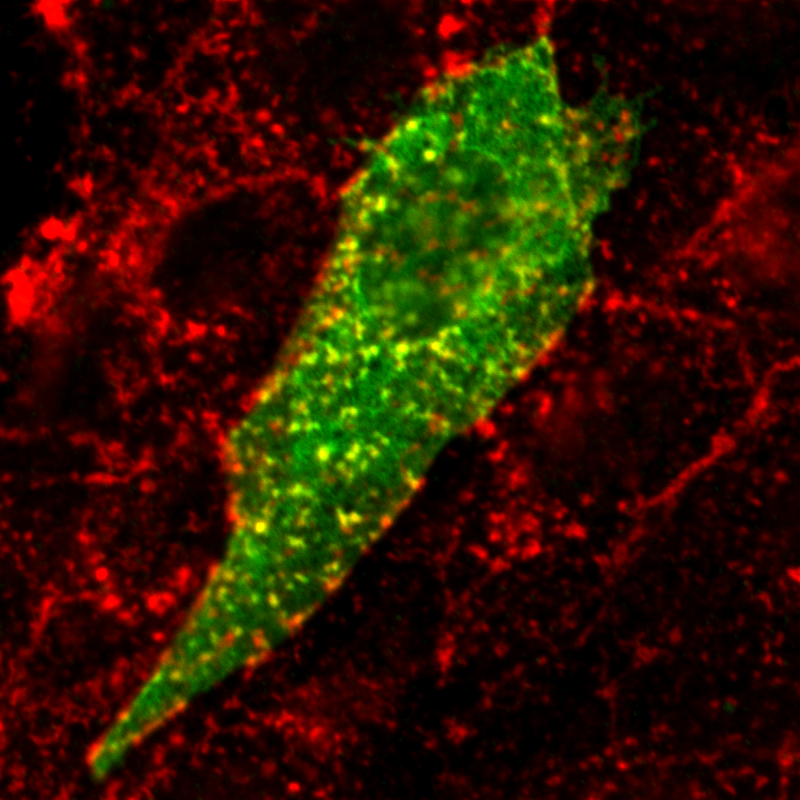
| organism | orgs |
|---|---|
| Organism: | Homo sapiens |